Expressions and Functions of Flowering Locus Ts in Narcissus tazetta var. chinensis Roem
-
摘要:
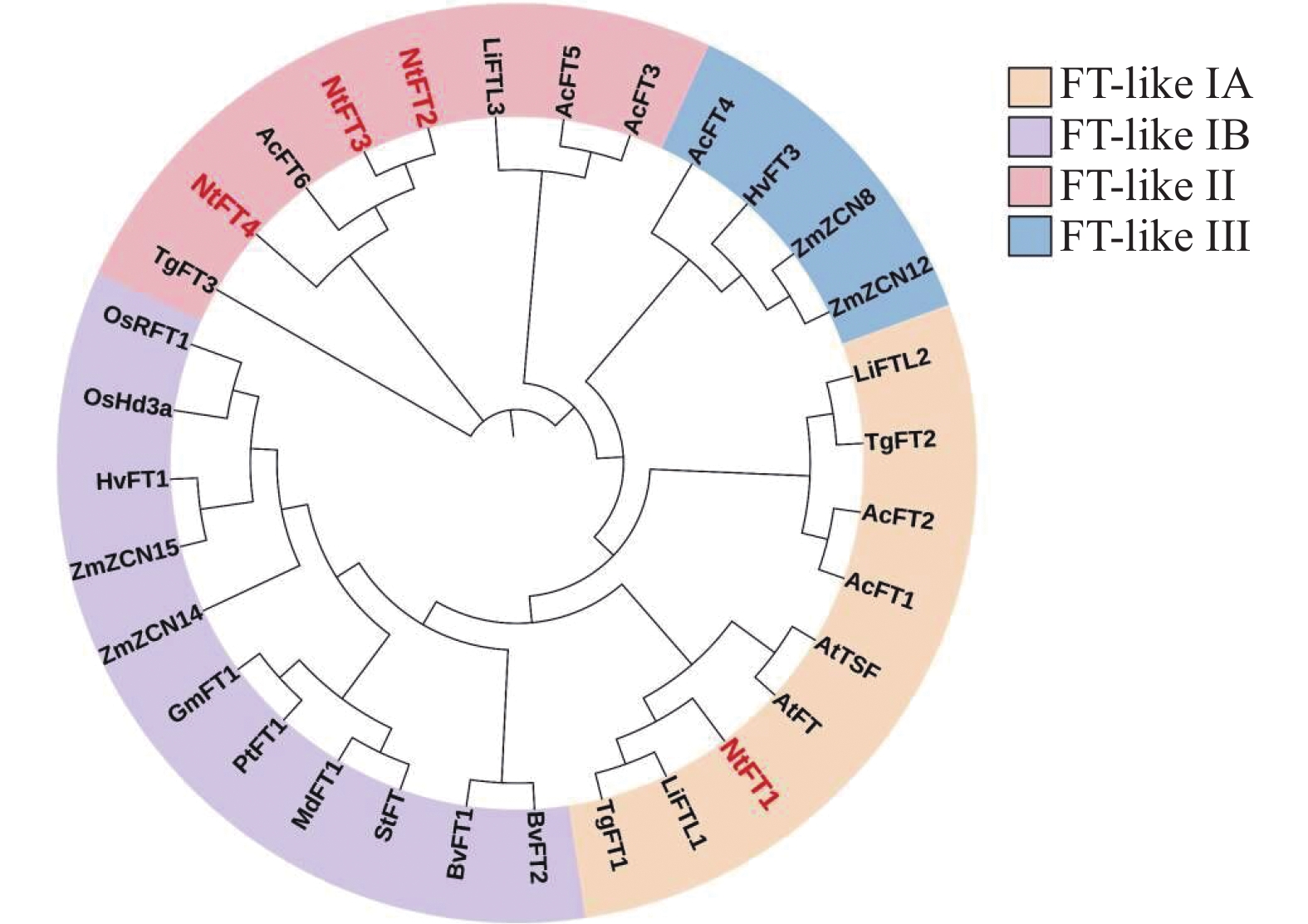
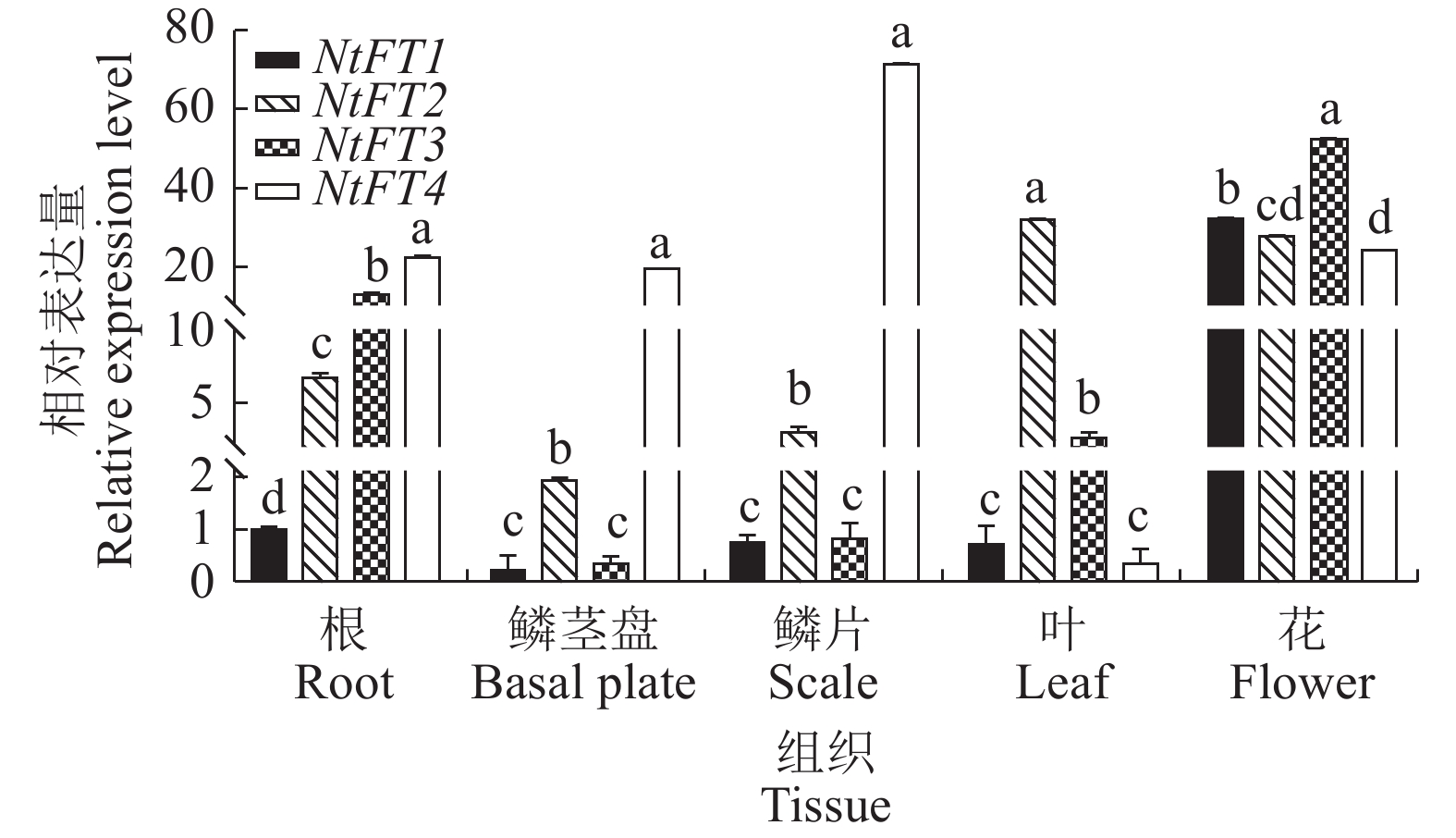
目的 开花基因座T(Flowering Locus T,FT)基因广泛参与植物的生长发育,在调控开花、地下茎发育、种子萌发和逆境胁迫等过程中发挥重要作用。揭示FT基因在中国水仙(Narcissus tazetta var. chinensis Roem)中的表达模式和功能可为中国水仙花期调控提供理论依据。 方法 基于中国水仙转录组数据,对FT 基因进行筛选,获得4个中国水仙FT基因。利用荧光定量PCR技术分析它们在中国水仙不同组织和花芽分化不同时期的表达模式,并在拟南芥(Arabidopsis thaliana)中超量表达。利用荧光定量PCR技术分析转NtFT1和NtFT4基因拟南芥中SUPPRESSOR OF OVEREXPRESSION OF CO 1(SOC1)、LEAFY(LFY)和APETALA 1(AP1) 基因表达水平。 结果 克隆了4个中国水仙FT同源基因NtFT1、NtFT2、NtFT3和NtFT4,除NtFT3外,中国水仙NtFT都具有FT的保守基序;系统进化分析显示,NtFT1属于FT-like I进化枝,NtFT2、NtFT3和NtFT4同属于FT-like II类进化枝。不同的NtFT基因在中国水仙组织器官和花芽分化不同时期的表达模式存在差异:NtFT1和NtFT3在花中表达量最高,NtFT2在叶片中表达量最高,NtFT4在鳞片中的表达量最高;NtFT1在花芽分化过程中均呈先上升后下降的趋势,NtFT2整个主芽分化过程中变化幅度不大,NtFT3和NtFT4在整个花芽分化期表达量都较低。异位转化拟南芥结果显示,与野生型拟南芥相比,过表达NtFT1和NtFT2的拟南芥提早开花,过表达NtFT3拟南芥开花时间与野生型植株无明显差异,转NtFT4基因的拟南芥植株推迟开花。过表达NtFT1拟南芥中SOC1、LFY和AP1基因表达量上升。 结论 在中国水仙中存在多个FT基因,在调控开花的功能上存在差异,NtFT1促进开花,NtFT4抑制开花。 Abstract:Objective Expressions and functions of Flowering Locus T (FT), the genes widely involved in plant growth, flowering regulation, root development, and seed germination, in Chinese narcissus were studied. Method Four FTs were identified from the transcriptome data on Narcissus tazetta var. chinensis Roem by bioinformatics analysis. Expressions of the genes in various tissues and flower buds at differentiation stages were determined by fluorescence quantitative PCR and further confirmed by overexpressing it in Arabidopsis thaliana. Result The 4 homologous FTs, i.e., NtFT1, NtFT2, NtFT3, and NtFT4, were cloned using RT-PCR. All of them, except NtFT3, had conserved motifs. Phylogenetically; NtFT1 belonged to the FT-like I branch, while NtFT2, NtFT3, and NtFT4, to the FT-like II branch. In tissues, organs, and flower buds at differentiation stages, they expressed differently. The highest expressions of NtFT1 and NtFT3 were in the flowers, those of NtFT2 in the leaves, and NtFT4 in the scales. At different flower bud differentiation stages, the expressions differed significantly as well, as NtFT1 decreased followed as an increase during differentiation, but NtFT2 changed little throughout. The expressions of NtFT3 and NtFT4 were relatively low in the entire flower bud differentiation period. The ectopic transformation of A. thaliana showed the overexpressed NtFT1 and NtFT2 led to early flowering in comparison to the wild type. But no effect was associated with NtFT3, but delayed flowering was found on the transgenic NtFT4 Arabidopsis plants. In addition, the expressions of SOC1, LFY, and AP1 in the Arabidopsis plants rose with NtFT1 overexpression. Conclusion There were multiple FTs in Chinese narcissus that differed in regulating the flowering—NtFT1 promoted, while NtFT4 inhibited, the process. -
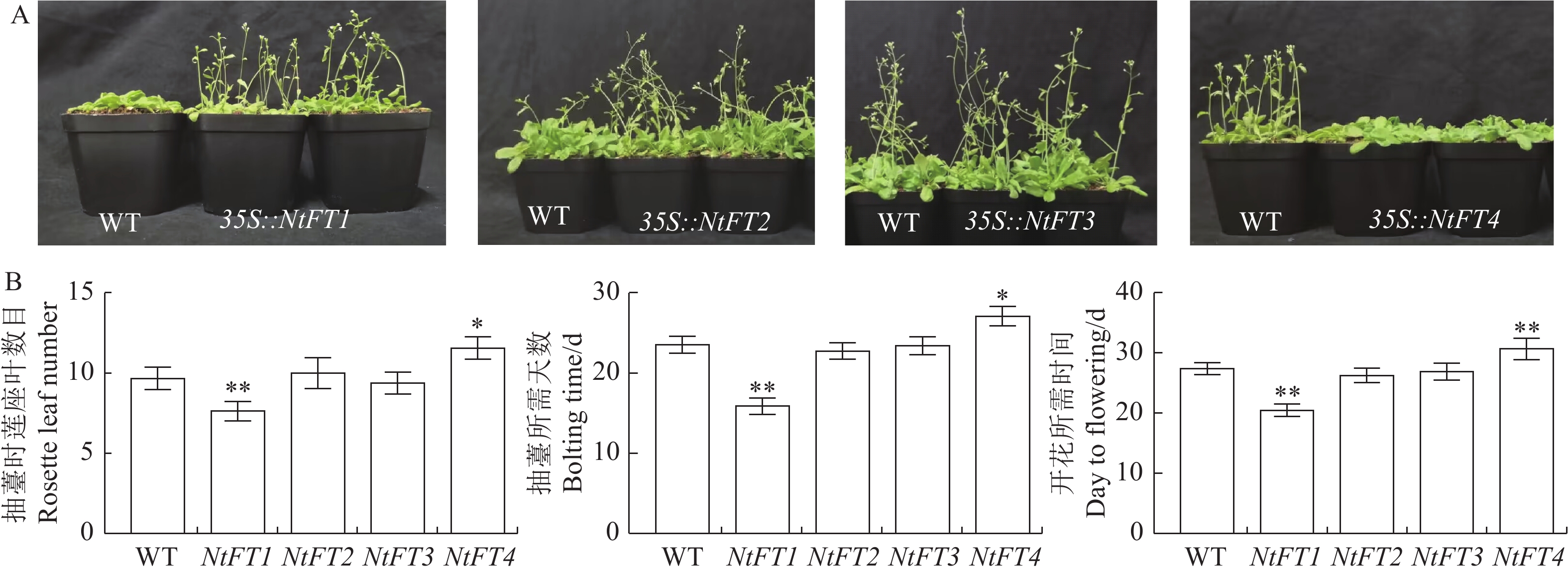
图 5 NtFT基因在拟南芥的异位表达
A:过表达NtFT基因拟南芥开花情况;B:过表达 NtFT 基因拟南芥开花性状数据。*、**表示与WT相比差异显著(P<0.05)或极显著(P<0.01)。图6同。
Figure 5. Ectopic expressions of NtFTs in A. thaliana
A: Flowering of A. thaliana with overexpressed NtFT; B: Flowering traits data of A. thaliana with overexpressed NtFT; * and ** indicate significant difference(P<0.05) and (P<0.01), respectively. Same for Fig. 6.
表 1 克隆中国水仙NtFT基因引物
Table 1. Primers for cloning NtFT of N. tazetta
引物名称
Primer name上游引物序列(5′-3′)
Upstream primer sequence (5′-3′)下游引物序列(5′-3′)
Downstream primer sequence (5′-3′)NtFT1 TTTCCGCTTATATCTCTTCTGGGAC TCGGGAAGTAGCAAGACGATCAAAC NtFT2 ATGTTGAGAGAGAGGGTACC TCAGCAAAGTCCTGAGAACCTTCTT NtFT3 GAAGTAGTCATGTTGAGAGAGAGG GTATCACATATTGCATGGCTTAGG NtFT4 GGTTAAGAGACAGAATGCCGAT TTTATGTCATTTATCGTCTG CTAG 表 2 FT基因表达qRT-PCR引物
Table 2. Primers for qRT-PCR of FT gene
引物名称
Primer name上游引物序列(5’-3’)
Upstream primer sequence (5′-3′)上游引物序列(5’-3’)
Downstream primer sequence (5′-3′)NtActin GTTGACCCACCACTAAGAACAATG TGCCCAGAAGTGCTATTCCAG QNtFT1 CCAGCCAAAGGTTGAAGTCG CCCTGTGGTTCCTGGTATG QNtFT2 CTTATGAGAGCCCTCGAACACC CGCACACAGTTTGTTGAACTTCC QNtFT3 CGCACACAGTTTGTTGAACTTCC CACTGGTTGGTGACAGACATACC QNtFT4 TGGCAGGATGCGATGCAAGA CACGATGCGATGAATCCCCGA 表 3 pSAK277表达载体构建引物
Table 3. Primers for constructing pSAK277 expression vector
引物名称
Primer name上游引物序列(5′-3′)
Upstream primer sequence (5′-3′)下游引物序列(5′-3′)
Upstream primer sequence (5′-3′)277NtFT1 ACTAGTGGATCCAAAGAATTCATGA
GTAGGGATCCTTTGGTTATTGAGAAGTACTCTCGAGAAGCTT
TTAGGGGTACATCCTCCGGCCACCA277NtFT2 ACTAGTGGATCCAAAGAATTCAGTTGA
GAGAGAGGGTACCAAGGGAGAAGTACTCTCGAGAAGCTTT
CAGCAAAGTCCTGAGAACCTTCTT277NtFT3 ACTAGTGGATCCAAAGAATTC
ATGTTGAGAG AGAGGGTACCAGAAGTACTCTCGAGAAGCTT
TCACATATTG CATGGCTTAG277NtFT4 ACTAGTGGATCCAAAGAATTCAT
GCCGATACTGGGACAAGTAGAAGTACTCTCGAGAAGCTTTC
AGAACCTTCTTCCTCCGCA表 4 转基因拟南芥的不同基因qRT-PCR引物
Table 4. Primers for qRT-PCR of different genes in transgenic A. thaliana
引物名称
Primer name上游引物序列(5’-3’)
Upstream primer sequence (5′-3′)上游引物序列(5’-3’)
Upstream primer sequence (5′-3′)Actin GCTGAGAGTTGATGGTGTGCT GGATACCCTTTCGCAGATAGAG AtLFY GTTAGGTTTTACGGCGAGCA GCAATCGTCTCCGTTCAGC AtAP1 ACCAAATCCAGCATCCTTACA TCAAGAGTCAGTTCGAGATCATTC AtSOC1 CTCTCAGTGCTTTGTGATGCT CGATTGAGCATGTTCCTATGCC -
[1] FORNARA F, DE MONTAIGU A, COUPLAND G. SnapShot: Control of flowering in Arabidopsis [J]. Cell, 2010, 141(3): 550−550.e2. doi: 10.1016/j.cell.2010.04.024 [2] 张艺能, 周玉萍, 陈琼华, 等. 拟南芥开花时间调控的分子基础 [J]. 植物学报, 2014, 49(4):469−482. doi: 10.3724/SP.J.1259.2014.00469ZHANG Y N, ZHOU Y P, CHEN Q H, et al. Molecular basis of flowering time regulation in Arabidopsis [J]. Chinese Bulletin of Botany, 2014, 49(4): 469−482.(in Chinese) doi: 10.3724/SP.J.1259.2014.00469 [3] KARLGREN A, GYLLENSTRAND N, KÄLLMAN T, et al. Evolution of the PEBP gene family in plants: Functional diversification in seed plant evolution [J]. Plant Physiology, 2011, 156(4): 1967−1977. doi: 10.1104/pp.111.176206 [4] BANFIELD M J, BARKER J J, PERRY A C, et al. Function from structure? The crystal structure of human phosphatidylethanolamine-binding protein suggests a role in membrane signal transduction [J]. Structure, 1998, 6(10): 1245−1254. doi: 10.1016/S0969-2126(98)00125-7 [5] VALVERDE F, MOURADOV A, SOPPE W, et al. Photoreceptor regulation of CONSTANS protein in photoperiodic flowering [J]. Science, 2004, 303(5660): 1003−1006. doi: 10.1126/science.1091761 [6] 张乔松, 杨伟儿. 中国水仙花芽分化及贮藏期外界因子对花序数的影响 [J]. 园艺学报, 1987, 14(2):139−143,145.ZHANG Q S, YANG W E. On flower-bud differentiation of Chinese narcissus and the effect of external factors in storage on flower percentage [J]. Acta Horticulturae Sinica, 1987, 14(2): 139−143,145.(in Chinese) [7] 申艳红, 姜涛, 赵湾湾, 等. 乙烯处理水仙催多花技术和机理的研究 [J]. 农业生物技术学报, 2019, 27(6):1003−1015.SHEN Y H, JIANG T, ZHAO W W, et al. Study on technology and mechanism of ethylene treatment promotes the formation of more flowers of Narcissus tazetta var. chinensis [J]. Journal of Agricultural Biotechnology, 2019, 27(6): 1003−1015.(in Chinese) [8] ODA A, NARUMI T, LI T P, et al. CsFTL3, a chrysanthemum FLOWERING LOCUS T-like gene, is a key regulator of photoperiodic flowering in chrysanthemums [J]. Journal of Experimental Botany, 2012, 63(3): 1461−1477. doi: 10.1093/jxb/err387 [9] MAO Y C, SUN J, CAO P P, et al. Functional analysis of alternative splicing of the FLOWERING LOCUS T orthologous gene in Chrysanthemum morifolium [J]. Horticulture Research, 2016, 3: 16058. doi: 10.1038/hortres.2016.58 [10] WANG L J, SUN J, REN L P, et al. CmBBX8 accelerates flowering by targeting CmFTL1 directly in summer chrysanthemum [J]. Plant Biotechnology Journal, 2020, 18(7): 1562−1572. doi: 10.1111/pbi.13322 [11] SUN J, WANG H, REN L P, et al. CmFTL2 is involved in the photoperiod- and sucrose-mediated control of flowering time in chrysanthemum [J]. Horticulture Research, 2017, 4: 17001. doi: 10.1038/hortres.2017.1 [12] OTAGAKI S, OGAWA Y, HIBRAND-SAINT OYANT L, et al. Genotype of FLOWERING LOCUS T homologue contributes to flowering time differences in wild and cultivated roses [J]. Plant Biology, 2015, 17(4): 808−815. doi: 10.1111/plb.12299 [13] CHEN L, CAI Y P, QU M N, et al. Soybean adaption to high-latitude regions is associated with natural variations of GmFT2b, an ortholog of FLOWERING LOCUS T [J]. Plant, Cell & Environment, 2020, 43(4): 934−944. [14] WU L, LI F, DENG Q H, et al. Identification and characterization of the FLOWERING LOCUS T/terminal flower 1 gene family in petunia [J]. DNA and Cell Biology, 2019, 38(9): 982−995. doi: 10.1089/dna.2019.4720 [15] HELLER W P, YING Z T, DAVENPORT T L, et al. Identification of members of the Dimocarpus longan flowering locus T gene family with divergent functions in flowering [J]. Tropical Plant Biology, 2014, 7(1): 19−29. doi: 10.1007/s12042-013-9134-0 [16] COELHO C P, MINOW M A A, CHALFUN-JÚNIOR A, et al. Putative sugarcane FT/TFL1 genes delay flowering time and alter reproductive architecture in Arabidopsis [J]. Frontiers in Plant Science, 2014, 5: 221. [17] NAVARRO C, ABELENDA J A, CRUZ-ORÓ E, et al. Control of flowering and storage organ formation in potato by FLOWERING LOCUS T [J]. Nature, 2011, 478(7367): 119−122. doi: 10.1038/nature10431 [18] NIWA M, DAIMON Y, KUROTANI K I, et al. BRANCHED1 interacts with FLOWERING LOCUS T to repress the floral transition of the axillary meristems in Arabidopsis [J]. The Plant Cell, 2013, 25(4): 1228−1242. doi: 10.1105/tpc.112.109090 [19] KINOSHITA T, ONO N, HAYASHI Y, et al. FLOWERING LOCUS T regulates stomatal opening [J]. Current Biology:CB, 2011, 21(14): 1232−1238. doi: 10.1016/j.cub.2011.06.025 [20] CHEN M, PENFIELD S. Feedback regulation of COOLAIR expression controls seed dormancy and flowering time [J]. Science, 2018, 360(6392): 1014−1017. doi: 10.1126/science.aar7361 [21] ANDRÉ D, MARCON A, LEE K C, et al. FLOWERING LOCUS T paralogs control the annual growth cycle in Populus trees [J]. Current Biology:CB, 2022, 32(13): 2988−2996.e4. doi: 10.1016/j.cub.2022.05.023 [22] 陈洁. 水稻FT-Like基因OsFTL4的功能研究[D]. 扬州: 扬州大学, 2020.CHEN J. Function research of FT-like gene OsTTL4 in rice [D]. Yangzhou: Yangzhou University, 2020. (in Chinese) [23] FENG Y, ZHU L Y, PAN T F, et al. Characterization of summer dormancy in Narcissus tazetta var. Chinensis and the role of NtFTs in summer dormancy and flower differentiation [J]. Scientia Horticulturae, 2015, 183: 109−117. doi: 10.1016/j.scienta.2014.11.013 [24] LI X F, JIA L Y, XU J, et al. FT-like NFT1 gene may play a role in flower transition induced by heat accumulation in Narcissus tazetta var. chinensis [J]. Plant and Cell Physiology, 2013, 54(2): 270−281. doi: 10.1093/pcp/pcs181 [25] CONANT G C, WOLFE K H. Turning a hobby into a job: How duplicated genes find new functions [J]. Nature Reviews Genetics, 2008, 9(12): 938−950. doi: 10.1038/nrg2482 [26] 李永光, 金玉环, 郭力, 等. 小鼠耳芥PEBP基因家族全基因组鉴定及表达分析 [J]. 遗传, 2022, 44(1):80−94.LI Y G, JIN Y H, GUO L, et al. Genome-wide identification and expression analysis of the PEBP genes in Arabidopsis pumila [J]. Hereditas(Beijing), 2022, 44(1): 80−94.(in Chinese) [27] 牛西强, 罗潇云, 康凯程, 等. 辣椒PEBP基因家族的全基因组鉴定、比较进化与组织表达分析 [J]. 园艺学报, 2021, 48(5):947−959.NIU X Q, LUO X Y, KANG K C, et al. Genome-wide identification, comparative evolution and expression analysis of PEBP gene family from Capsicum annuum [J]. Acta Horticulturae Sinica, 2021, 48(5): 947−959.(in Chinese) [28] JIANG X D, ZHONG M C, DONG X, et al. Rosoideae-specific duplication and functional diversification of FT-like genes in Rosaceae [J]. Horticulture Research, 2022, 9: uhac059. doi: 10.1093/hr/uhac059 [29] LIU H L, LIU X, CHANG X J, et al. Large-scale analyses of angiosperm Flowering Locus T genes reveal duplication and functional divergence in monocots [J]. Frontiers in Plant Science, 2023, 13: 1039500. doi: 10.3389/fpls.2022.1039500 [30] LEEGGANGERS H A, ROSILIO-BRAMI T, BIGAS-NADAL J, et al. Tulipa gesneriana and Lilium longiflorum PEBP genes and their putative roles in flowering time control [J]. Plant and Cell Physiology, 2018, 59(1): 90−106. doi: 10.1093/pcp/pcx164 [31] LEE R, BALDWIN S, KENEL F, et al. FLOWERING LOCUS T genes control onion bulb formation and flowering [J]. Nature Communications, 2013, 4: 2884. doi: 10.1038/ncomms3884 [32] YAN X, CAO Q Z, HE H B, et al. Functional analysis and expression patterns of members of the FLOWERING LOCUS T (FT) gene family in Lilium [J]. Plant Physiology and Biochemistry, 2021, 163: 250−260. doi: 10.1016/j.plaphy.2021.03.056 [33] KOTODA N, HAYASHI H, SUZUKI M, et al. Molecular characterization of FLOWERING LOCUS T-like genes of apple (Malus ×domestica borkh. ) [J]. Plant and Cell Physiology, 2010, 51(4): 561−575. doi: 10.1093/pcp/pcq021 [34] 朱燕宇. 小叶杨FT基因家族的克隆及功能验证[D]. 南京: 南京林业大学, 2015.ZHU Y Y. Cloning and functional analysis of FT gene family from Populus simonii[D]. Nanjing: Nanjing Forestry University, 2015. (in Chinese) [35] PIN P A, NILSSON O. The multifaceted roles of FLOWERING LOCUS T in plant development [J]. Plant, Cell & Environment, 2012, 35(10): 1742−1755. [36] LIU W, JIANG B J, MA L M, et al. Functional diversification of Flowering Locus T homologs in soybean: GmFT1a and GmFT2a/5a have opposite roles in controlling flowering and maturation [J]. The New Phytologist, 2018, 217(3): 1335−1345. doi: 10.1111/nph.14884 -







 下载:
下载: