Sequence and Annotation of Colletotrichum fructicola N425 Genome on Tea Plant
-
摘要:
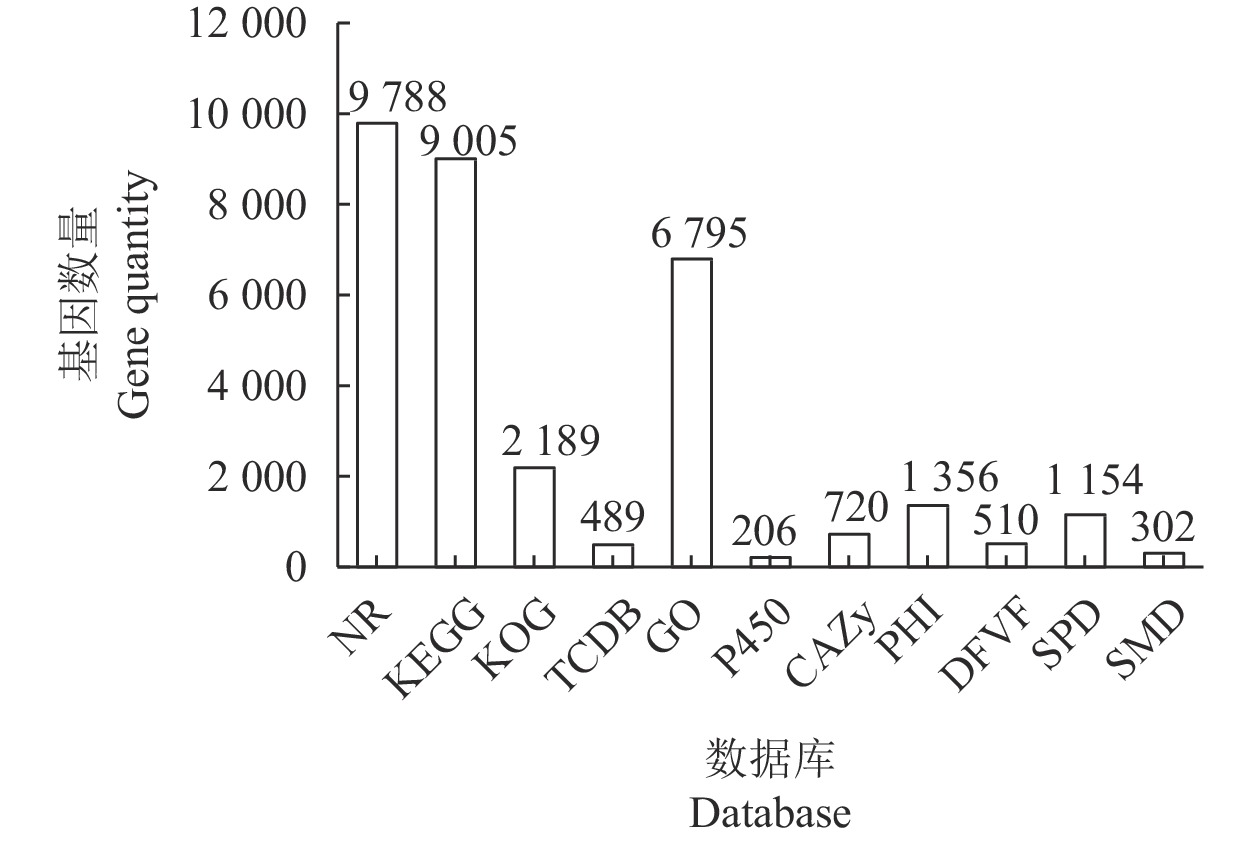
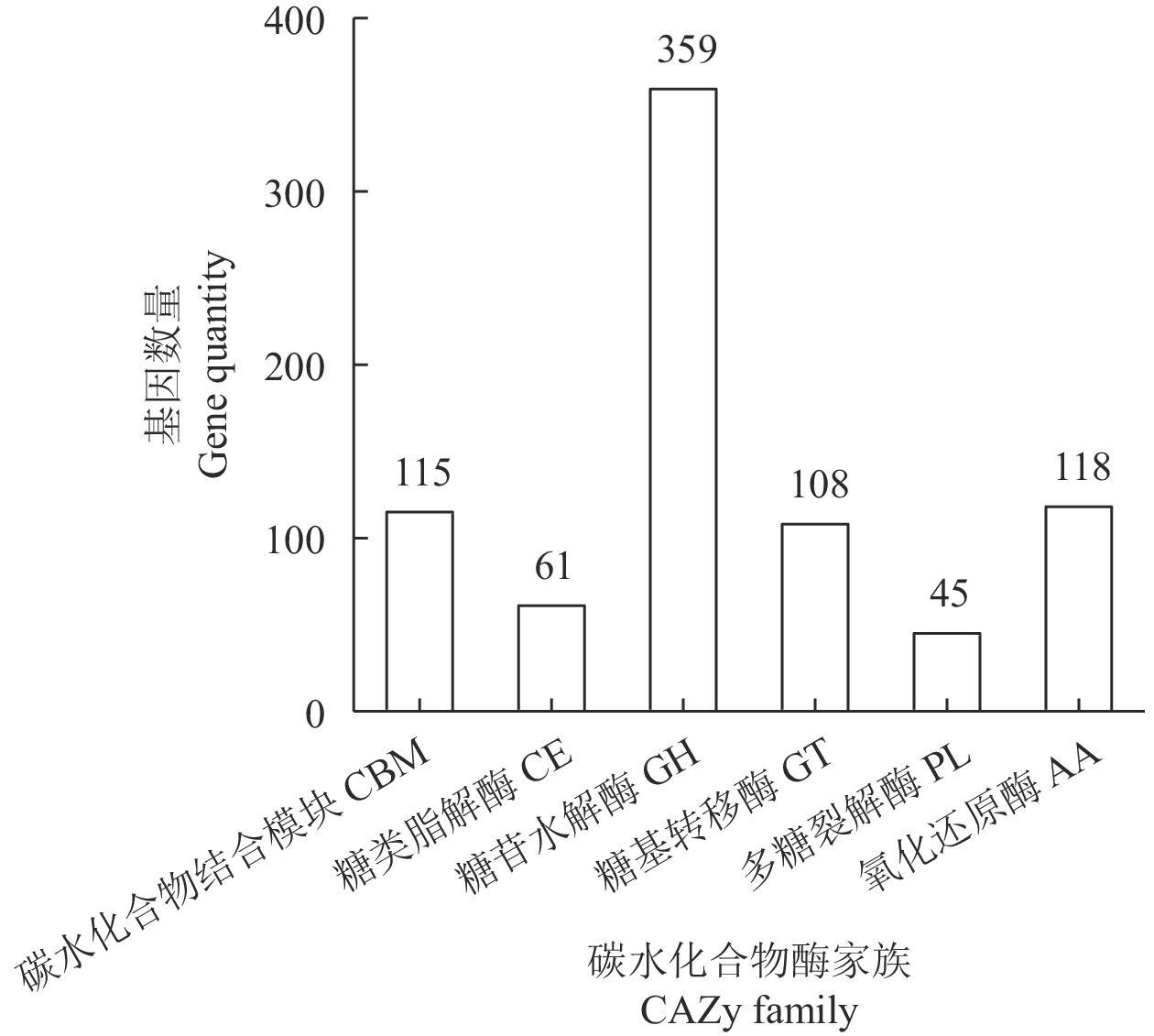
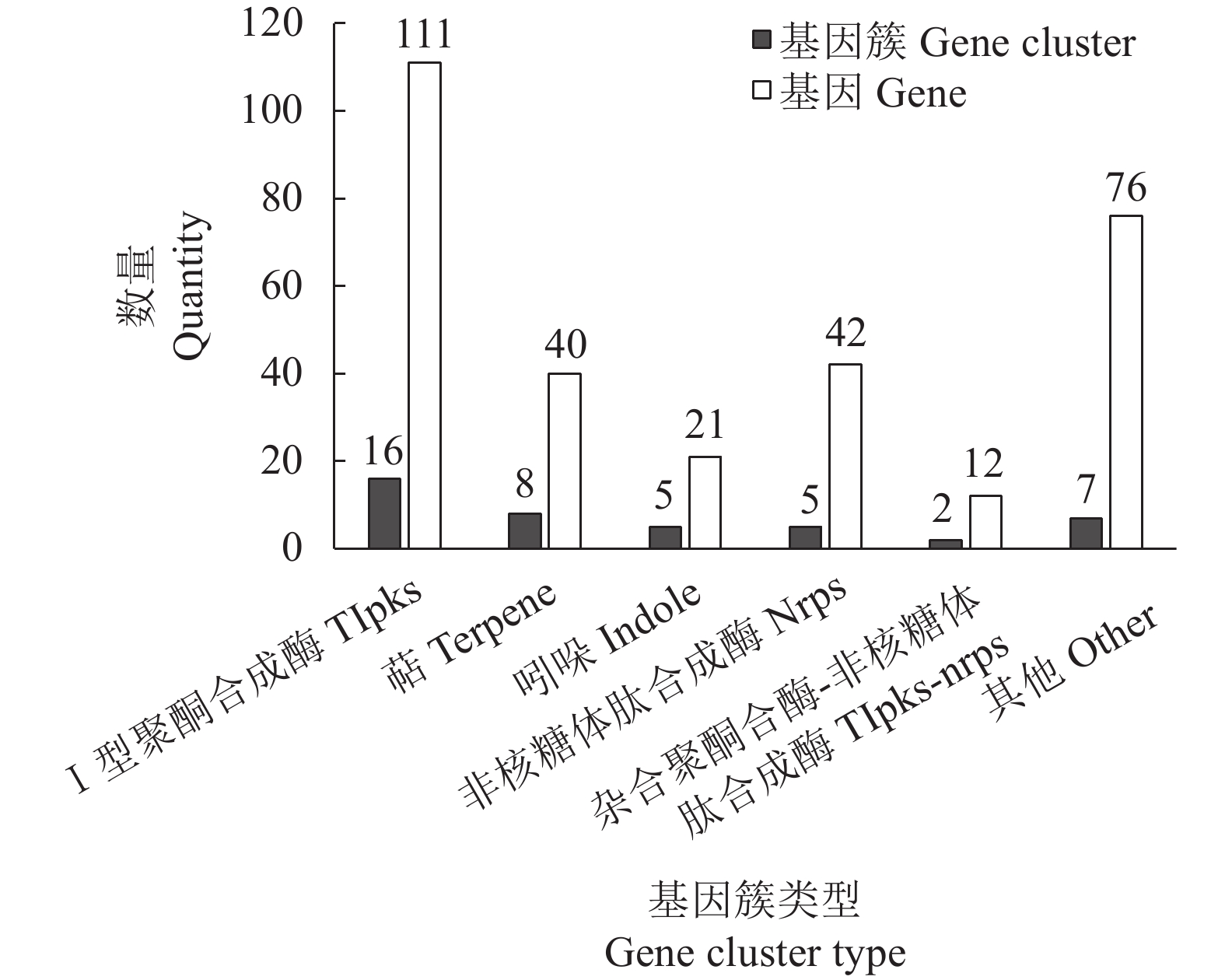
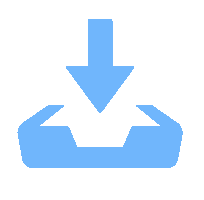
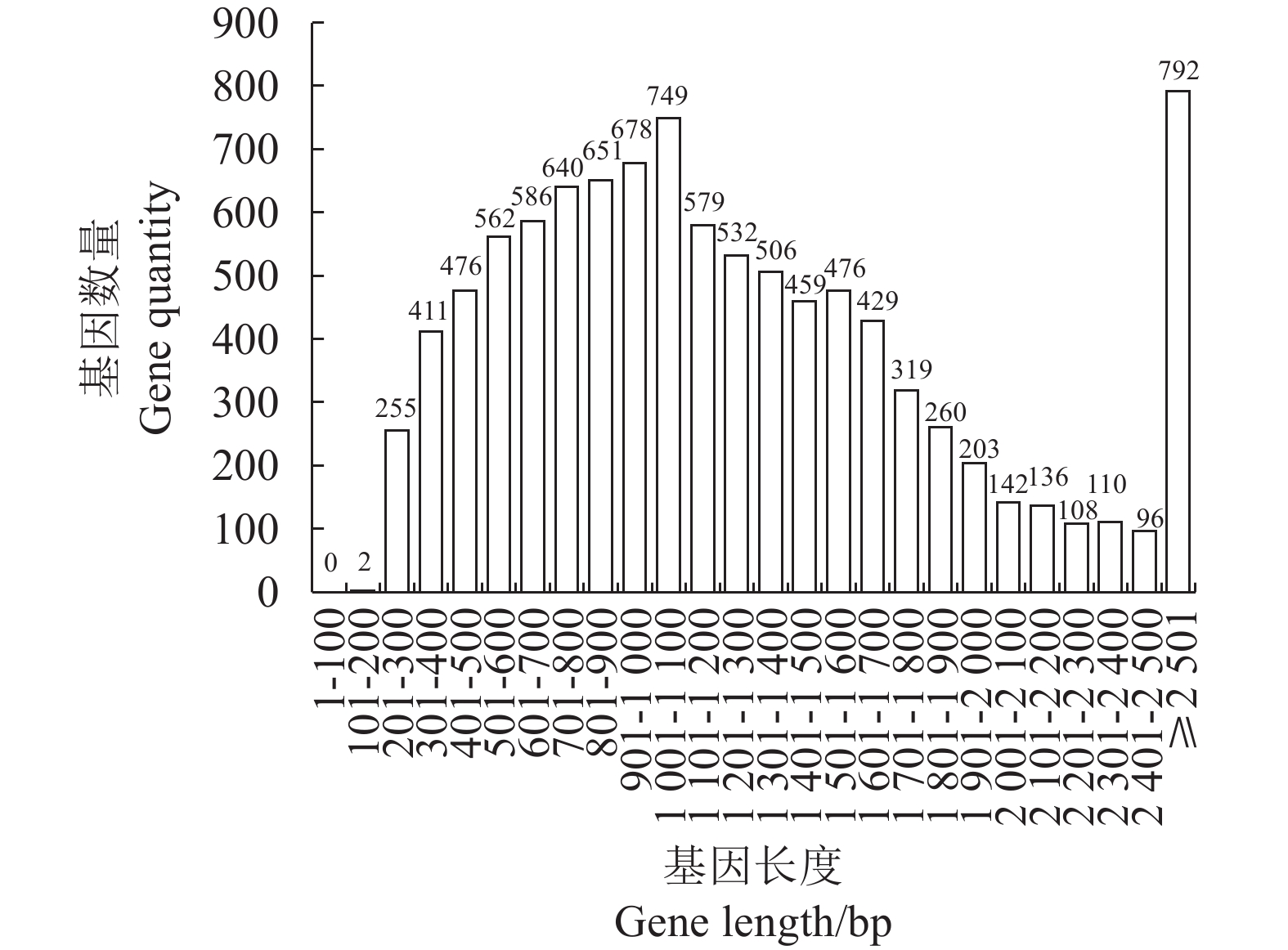
目的 对茶树炭疽菌N425进行全基因组测序和基因功能注释并从中初筛出致病相关基因。 方法 通过Illumina HiSeq2500 PE150平台对N425菌株进行全基因组测序和组装,并利用NR、KEGG、KOG等功能基因数据库进行基因预测和功能注释。 结果 茶树炭疽菌N425基因组全长约56.3 Mbp,G+C含量为53.2%,共预测出10 157个蛋白编码基因和若干各类非蛋白编码序列。其中,1 356个基因在病原与宿主互作数据库(PHI)中得到注释,包括140个注释为致病力丧失的基因;720个基因在碳水化合物酶数据库(CAZy)中得到注释;302个基因被预测为次生代谢物相关基因。这些基因中有涉及cAMP-PKA、MAPK等信号通路以及降解宿主细胞壁等致病相关过程,是潜在的致病相关基因。 结论 本研究成功获得茶树炭疽菌N425全基因组序列和基因功能注释,并从中挖掘出潜在致病相关基因,为后续展开对茶树炭疽菌侵染茶树的致病分子机制研究奠定基础。 Abstract:Objective Sequence and annotation of Colletotrichum fructicola N425 genome from a diseased tea plant were determined, and primary virulence-related genes identified. Method Whole genome of N425 was sequenced and assembled using the Illumina HiSeq2500 PE150 platform. Predicted protein structure and functional annotation of the genes were obtained using the NR, KEGG, and KOG databases. Results The genome was approximately 56.3 Mbp in length with 53.2% of G+C, 10 157 protein-coding genes, and several types of non-protein coding sequences. Of the genes, 1 356 were annotated in the PHI database that included 140 as the loss of pathogenicity, 720 in the CAZy database, and 302 related to the secondary metabolites. Some of these genes were involved in the cAMP-PKA and MAPK cascade signaling pathways as well as other pathogenic processes, such as host cell wall degradation, which could be responsible for the virulence on tea plants. Conclusion The sequence and annotation of whole C. fructicola N425 genome were successfully obtained, and probable virulence-related genes identified. -
Key words:
- Tea plant anthracnose /
- whole-genome sequence /
- gene annotation /
- virulence-related genes
-
表 1 经PHI数据库归类为致病性丧失的前18个基因的注释信息
Table 1. Annotation on first 18 loss of pathogenicity genes classified by PHI database
基因编号
Gene No.同源基因名称
Homologous gene name同源基因所属病原菌
Pathogen to which homologous gene belongs宿主
Host基因注释
Gene annotation参考文献
ReferenceA00871 ChMK1 希金斯炭疽菌
Colletotrichum higginsianum拟南芥
Thale cress促分裂原活化蛋白激酶
Mitogen-activated protein kinase(MAPK)[17] A09504 CgAP1 胶孢炭疽菌
C. gloeosporioides芒果
MangobZIP 转录因子
bZIP transcription factor[18] A08060 CAP20 胶孢炭疽菌
C. gloeosporioides牛油果
Avocado附着胞细胞壁
Appressorial wall[19] A03691 CAM 稻瘟病菌
Magnaporthe oryzae小麦
Wheat钙调蛋白
Calmodulin[20] A04430 MoYpt7 稻瘟病菌
M. oryzae水稻
Rice小GTP酶
Small GTPase[21] A04622 ClaSSD1 葫芦炭疽菌
C. lagenaria黄瓜
Cucumber细胞壁组装调控
Regulation of cell wall assembly[22] A09136 ChSte7 希金斯炭疽菌
C. higginsianum拟南芥
Thale cress促分裂原活化蛋白激酶
Mitogen-activated protein kinase kinase(MAPKK)[23] A01252 RAC1 稻瘟病菌
M. oryzae水稻
RiceRhoGTP酶
Rho GTPase[24] A00884 CNB 稻瘟病菌
M. oryzae小麦
Wheat钙调磷酸酶调节亚基
Calcineurin regulatory subunits[20] A03506 CLAP1 菜豆炭疽病菌
C. lindemuthianum四季豆
String bean铜转运ATP酶
Copper transporting ATPase[25] A02922 ChPMA2 希金斯炭疽菌
C. higginsianum拟南芥
Thale cress质膜质子泵
Plasma membrane proton pump[26] A09008 GPI8 禾生炭疽菌
C. graminicola玉米
Maize半胱氨酸蛋白酶
Cystein protease[27] A06226 CgPKS1 禾生炭疽菌
C. graminicola玉米
Maize聚酮合酶
Polyketide synthase[28] A02402 ACL 禾谷镰刀菌
Fusarium graminearum小麦
WheatATP柠檬酸裂解酶
ATP citrate lyase[29] A09126 RPK1 葫芦炭疽菌
C. lagenaria黄瓜
Cucumber蛋白激酶A调节亚基
Protein kinase A(PKA)regulatory subunit[30] A05844 FGA2 尖孢镰刀菌
F. oxysporum西红柿
TomatoG蛋白α亚基
G protein alpha subunit[31] A01313 MgRho3 稻瘟病菌
M. oryzae水稻
RiceRhoGTP酶
Rho GTPase[32] A07576 VdNoxB 大丽轮枝菌
Verticillium dahliae棉花
CottonNADPH 氧化酶
NADPH oxidase[33] 表 2 部分CAZy基因的注释信息
Table 2. Annotation on some CAZy genes
基因编号
Gene No.同源基因GenBank登录号
Homologous gene
GenBank accession No.同源基因所属病原菌
Pathogen to which homologous gene belongs基因注释
Gene annotationCAZy_家族
CAZy_familyA02429 AUT30986.1 胶孢炭疽菌 Colletotrichum gloeosporioides 果胶裂解酶2 Pectin lyase 2 PL1_4 A03941 ELA25906.1 果生炭疽菌 C. fructicola 半乳糖氧化酶前体 Galactose oxidase precursor AA5_2 A03990 AUT30987.1 胶孢炭疽菌 C. gloeosporioides 多聚半乳糖醛酸内切酶1 Endopolygalacturonase 1 GH28 A07825 AUT30984.1 胶孢炭疽菌 C. gloeosporioides 果胶酸裂解酶B Pectate lyase B PL1_7 A00270 AHC30245.1 胶孢炭疽菌 C. gloeosporioides 漆酶2 Laccase 2 AA1_3 A00454 AND80748.1 胶孢炭疽菌 C. gloeosporioides 内切葡聚糖酶1B Endoglucanase 1B GH12 A07600 AAL38030.1 胶孢炭疽菌 C. gloeosporioides 角质酶 Cutinase CE5 A06012 AFK76451.1 胶孢炭疽菌 C. gloeosporioides 漆酶 Laccase AA1_3 A07644 AAD00894.3 胶孢炭疽菌 C. gloeosporioides 甾醇糖基转移酶 Sterol glycosyltransferase GT1 A07350 AUT30983.1 胶孢炭疽菌 C. gloeosporioides 果胶酸裂解酶A Pectate lyase A PL1_7 A04870 AUT30985.1 胶孢炭疽菌 C. gloeosporioides 果胶裂解酶1 Pectin lyase 1 CBM1/PL1_4 A02466 AAG32629.1 里氏木霉 Trichoderma reesei 甘露糖磷酸醇合成酶 Mannose phospho-dolichol synthase GT2 A04591 CAP73770.1 柄孢霉 Podospora anserina 糖原合成酶 Glycogen synthase GT3 A08316 AAL23719.1 禾生炭疽菌 C. graminicola 几丁质合成酶C Chitin synthase C GT2 A00674 AAL23718.1 禾生炭疽菌 C. graminicola 几丁质合成酶B Chitin synthase B GT2 A06213 AAL23717.1 禾生炭疽菌 C. graminicola 几丁质合成酶A Chitin synthase A GT2 -
[1] 刘威, 袁丁, 郭桂义, 等. 茶树炭疽病病原鉴定 [J]. 南方农业学报, 2017, 48(3):448−453.LIU W, YUAN D, GUO G Y, et al. Identification of anthracnose pathogen in tea plant [J]. Journal of Southern Agriculture, 2017, 48(3): 448−453.(in Chinese) [2] WANG Y C, HAO X Y, WANG L, et al. Diverse Colletotrichum species cause anthracnose of tea plants (Camellia sinensis (L. ) O. Kuntze) in China [J]. Scientific Reports, 2016, 6: 35287. doi: 10.1038/srep35287 [3] GUO M, PAN Y M, DAI Y L, et al. First report of brown blight disease caused by Colletotrichum gloeosporioides on Camellia sinensis in Anhui Province, China [J]. Plant Disease, 2014, 98(2): 284. [4] O'CONNELL R J, THON M R, HACQUARD S, et al. Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses [J]. Nature Genetics, 2012, 44(9): 1060−1065. doi: 10.1038/ng.2372 [5] DEAN R A, TALBOT N J, EBBOLE D J, et al. The genome sequence of the rice blast fungus Magnaporthe grisea [J]. Nature, 2005, 434(7036): 980−986. doi: 10.1038/nature03449 [6] CUOMO C A, GÜLDENER U, XU J R, et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization [J]. Science, 2007, 317(5843): 1400−1402. doi: 10.1126/science.1143708 [7] LIANG X F, WANG B, DONG Q Y, et al. Pathogenic adaptations of Colletotrichum fungi revealed by genome wide gene family evolutionary analyses [J]. PLoS One, 2018, 13(4): e0196303. doi: 10.1371/journal.pone.0196303 [8] LIANG X F, WEI T Y, CAO M Y, et al. The MAP kinase CfPMK1 is a key regulator of pathogenesis, development, and stress tolerance of Colletotrichum fructicola [J]. Frontiers in Microbiology, 2019, 10: 1070. doi: 10.3389/fmicb.2019.01070 [9] BI F C, MENT D, LURIA N, et al. Mutation of AREA affects growth, sporulation, nitrogen regulation, and pathogenicity in Colletotrichum gloeosporioides [J]. Fungal Genetics and Biology, 2017, 99: 29−39. doi: 10.1016/j.fgb.2016.12.006 [10] PAN Y T, LI L W, YANG J Y, et al. Involvement of protein kinase CgSat4 in potassium uptake, cation tolerance, and full virulence in Colletotrichum gloeosporioides [J]. Frontiers in Plant Science, 2022, 13: 773898. doi: 10.3389/fpls.2022.773898 [11] YANG G Y, YANG J, ZHANG Q W, et al. The effector protein CgNLP1 of Colletotrichum gloeosporioides affects invasion and disrupts nuclear localization of necrosis-induced transcription factor HbMYB8-like to suppress plant defense signaling [J]. Frontiers in Microbiology, 2022, 13: 911479. doi: 10.3389/fmicb.2022.911479 [12] GAN R C, ZHANG S P, LI H. Cell wall integrity mediated by CfCHS1 is important for growth, stress responses and pathogenicity in Colletotrichum fructicola [J]. Journal of Fungi, 2023, 9(6): 643. doi: 10.3390/jof9060643 [13] KONG Y Y, YUAN Y L, YANG M H, et al. CfCpmd1 regulates pathogenicity and sexual development of plus and minus strains in Colletotrichum fructicola causing Glomerella leaf spot on apple in China [J]. Phytopathology, 2023, 113(10): 1985−1993. [14] 彭成彬, 陈美霞, 魏日凤, 等. 茶树炭疽菌分离鉴定与遗传转化体系建立 [J]. 西南农业学报, 2021, 34(10):2167−2173.PENG C B, CHEN M X, WEI R F, et al. Isolation and identification of Anthrax from tea plant and establishment of genetic transformation system [J]. Southwest China Journal of Agricultural Sciences, 2021, 34(10): 2167−2173.(in Chinese) [15] XIA Y M, CHEN F S, DU Y, et al. A modified SDS-based DNA extraction method from raw soybean [J]. Bioscience Reports, 2019, 39(2): BSR20182271. doi: 10.1042/BSR20182271 [16] 顾鑫, 杨晓贺, 姚亮亮, 等. 大豆灰斑病菌Race15的全基因组测序分析 [J]. 大豆科学, 2021, 40(4):466−475.GU X, YANG X H, YAO L L, et al. Whole-genome sequencing and analysis of Cercospora sojina race 15 [J]. Soybean Science, 2021, 40(4): 466−475.(in Chinese) [17] WEI W, XIONG Y, ZHU W J, et al. Colletotrichum higginsianum mitogen-activated protein kinase ChMK1: Role in growth, cell wall integrity, colony melanization, and pathogenicity [J]. Frontiers in Microbiology, 2016, 7: 1212. [18] SUN Y J, WANG Y L, TIAN C M. bZIP transcription factor CgAP1 is essential for oxidative stress tolerance and full virulence of the poplar anthracnose fungus Colletotrichum gloeosporioides [J]. Fungal Genetics and Biology, 2016, 95: 58−66. doi: 10.1016/j.fgb.2016.08.006 [19] HWANG C S, FLAISHMAN M A, KOLATTUKUDY P E. Cloning of a gene expressed during appressorium formation by Colletotrichum gloeosporioides and a marked decrease in virulence by disruption of this gene [J]. The Plant Cell, 1995, 7(2): 183−193. [20] NGUYEN Q B, KADOTANI N, KASAHARA S, et al. Systematic functional analysis of calcium-signalling proteins in the genome of the rice-blast fungus, Magnaporthe oryzae, using a high-throughput RNA-silencing system [J]. Molecular Microbiology, 2008, 68(6): 1348−1365. doi: 10.1111/j.1365-2958.2008.06242.x [21] LIU X H, CHEN S M, GAO H M, et al. The small GTPase MoYpt7 is required for membrane fusion in autophagy and pathogenicity of Magnaporthe oryzae [J]. Environmental Microbiology, 2015, 17(11): 4495−4510. doi: 10.1111/1462-2920.12903 [22] TANAKA S, YAMADA K, YABUMOTO K, et al. Saccharomyces cerevisiae SSD1 orthologues are essential for host infection by the ascomycete plant pathogens Colletotrichum lagenarium and Magnaporthe grisea [J]. Molecular Microbiology, 2007, 64(5): 1332−1349. doi: 10.1111/j.1365-2958.2007.05742.x [23] YUAN Q F, CHEN M J, YAN Y Q, et al. ChSte7 is required for vegetative growth and various plant infection processes in Colletotrichum higginsianum[J]. BioMed Research International, 2016: 1−11. [24] CHEN J S, ZHENG W, ZHENG S Q, et al. Rac1 is required for pathogenicity and Chm1-dependent conidiogenesis in rice fungal pathogen Magnaporthe grisea [J]. PLoS Pathogens, 2008, 4(11): e1000202. doi: 10.1371/journal.ppat.1000202 [25] PARISOT D, DUFRESNE M, VENEAULT C, et al. clap1, a gene encoding a copper-transporting ATPase involved in the process of infection by the phytopathogenic fungus Colletotrichum lindemuthianum [J]. Molecular Genetics and Genomics, 2002, 268(2): 139−151. doi: 10.1007/s00438-002-0744-8 [26] KORN M, SCHMIDPETER J, DAHL M, et al. A genetic screen for pathogenicity genes in the hemibiotrophic fungus Colletotrichum higginsianum identifies the plasma membrane proton pump Pma2 required for host penetration [J]. PLoS One, 2015, 10(5): e0125960. doi: 10.1371/journal.pone.0125960 [27] OLIVEIRA-GARCIA E, DEISING H B. The glycosylphosphatidylinositol anchor biosynthesis genes GPI12, GAA1, and GPI8 are essential for cell-wall integrity and pathogenicity of the maize anthracnose fungus Colletotrichum graminicola [J]. Molecular Plant-Microbe Interactions, 2016, 29(11): 889−901. doi: 10.1094/MPMI-09-16-0175-R [28] LUDWIG N, LÖHRER M, HEMPEL M, et al. Melanin is not required for turgor generation but enhances cell-wall rigidity in appressoria of the corn pathogen Colletotrichum graminicola [J]. Molecular Plant-Microbe Interactions, 2014, 27(4): 315−327. doi: 10.1094/MPMI-09-13-0267-R [29] SON H, LEE J, PARK A R, et al. ATP citrate lyase is required for normal sexual and asexual development in Gibberella zeae [J]. Fungal Genetics and Biology, 2011, 48(4): 408−417. doi: 10.1016/j.fgb.2011.01.002 [30] TAKANO Y, KOMEDA K, KOJIMA K, et al. Proper regulation of cyclic AMP-dependent protein kinase is required for growth, conidiation, and appressorium function in the anthracnose fungus Colletotrichum lagenarium [J]. Molecular Plant-Microbe Interactions, 2001, 14(10): 1149−1157. doi: 10.1094/MPMI.2001.14.10.1149 [31] JAIN S, AKIYAMA K, TAKATA R, et al. Signaling via the G protein alpha subunit FGA2 is necessary for pathogenesis in Fusarium oxysporum [J]. FEMS Microbiology Letters, 2005, 243(1): 165−172. doi: 10.1016/j.femsle.2004.12.009 [32] ZHENG W, CHEN J S, LIU W D, et al. A Rho3 homolog is essential for appressorium development and pathogenicity of Magnaporthe grisea [J]. Eukaryotic Cell, 2007, 6(12): 2240−2250. doi: 10.1128/EC.00104-07 [33] ZHAO Y L, ZHOU T T, GUO H S. Hyphopodium-specific VdNoxB/VdPls1-dependent ROS-Ca2+ signaling is required for plant infection by Verticillium dahliae [J]. PLoS Pathogens, 2016, 12(7): e1005793. doi: 10.1371/journal.ppat.1005793 [34] LUO K, GUO J, HE D J, et al. Deoxynivalenol accumulation and detoxification in cereals and its potential role in wheat-Fusarium graminearum interactions [J]. aBIOTECH, 2023, 4(2): 155−171. doi: 10.1007/s42994-023-00096-7 [35] GUPTA L, VERMANI M, KAUR AHLUWALIA S, et al. Molecular virulence determinants of Magnaporthe oryzae: Disease pathogenesis and recent interventions for disease management in rice plant [J]. Mycology, 2021, 12(3): 174−187. doi: 10.1080/21501203.2020.1868594 [36] LI G T, ZHOU X Y, XU J R. Genetic control of infection-related development in Magnaporthe oryzae [J]. Current Opinion in Microbiology, 2012, 15(6): 678−684. doi: 10.1016/j.mib.2012.09.004 [37] ZHANG C K, WANG Y, WANG J Q, et al. Functional characterization of Rho family small GTPases in Fusarium graminearum [J]. Fungal Genetics and Biology, 2013, 61: 90−99. doi: 10.1016/j.fgb.2013.09.001 [38] KIM Y K, WANG Y H, LIU Z M, et al. Identification of a hard surface contact-induced gene in Colletotrichum gloeosporioides conidia as a sterol glycosyl transferase, a novel fungal virulence factor [J]. The Plant Journal, 2002, 30(2): 177−187. doi: 10.1046/j.1365-313X.2002.01284.x [39] SKAMNIOTI P, FURLONG R F, GURR S J. Evolutionary history of the ancient cutinase family in five filamentous Ascomycetes reveals differential gene duplications and losses and in Magnaporthe grisea shows evidence of sub- and neo-functionalization [J]. The New Phytologist, 2008, 180(3): 711−721. doi: 10.1111/j.1469-8137.2008.02598.x [40] SKAMNIOTI P, GURR S J. Magnaporthe grisea Cutinase2 mediates appressorium differentiation and host penetration and is required for full virulence [J]. The Plant Cell, 2007, 19(8): 2674−2689. doi: 10.1105/tpc.107.051219 -






 下载:
下载: